Deleted
Deleted Member
Posts: 0
|
Post by Deleted on Feb 29, 2020 13:29:41 GMT -5
|
|
|
|
Post by brobear on Oct 24, 2020 23:49:10 GMT -5
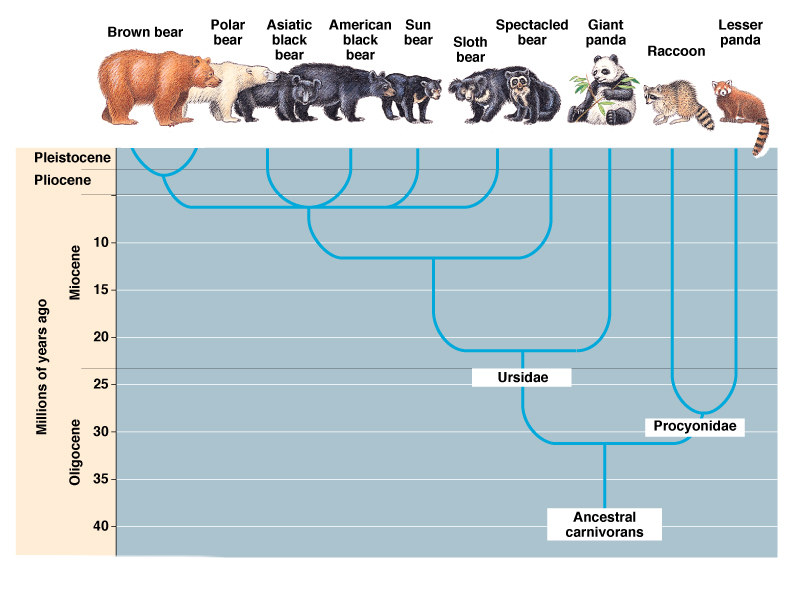
The Bear's Family Tree:

|
|
|
|
Post by brobear on Oct 24, 2020 23:50:16 GMT -5
Arctoidea is an infraorder of mostly carnivorous mammals which include the extinct Hemicyonidae (dog-bears), and the extant Musteloidea (weasels, raccoons, skunks, red pandas), Pinnipedia (seals, sea lions), and Ursidae (bears), found in all continents from the Eocene, 46 million years ago, to the present. Arctoids are caniforms, along with dogs (canids) and extinct bear dogs (Amphicyonidae). The earliest caniforms were superficially similar to martens, which are tree-dwelling mustelids. Together with feliforms, caniforms comprise the order Carnivora.
|
|
|
|
Post by brobear on Oct 25, 2020 17:00:41 GMT -5
Scientific classification:
Kingdom: Animalia
Phylum: Chordata
Class: Mammalia
Order: Carnivora
Infraorder: Arctoidea
Family: Ursidae
Subfamilies:
Amphicynodontinae
Hemicyoninae
Ursavinae
Agriotheriinae
Ailuropodinae
Tremarctinae
Ursinae
|
|
|
|
Post by brobear on Jan 5, 2021 5:02:15 GMT -5
This is a topic that completely leaves me in the dark, but a subject often mentioned throughout the Domain within numerous topics. So, I decided to post a topic page just for the subject - for those few who might have some understanding of it.  academic.oup.com/mbe/article/31/8/2004/2925840 academic.oup.com/mbe/article/31/8/2004/2925840 Bears in a Forest of Gene Trees: Phylogenetic Inference Is Complicated by Incomplete Lineage Sorting and Gene Flow. Abstract Ursine bears are a mammalian subfamily that comprises six morphologically and ecologically distinct extant species. Previous phylogenetic analyses of concatenated nuclear genes could not resolve all relationships among bears, and appeared to conflict with the mitochondrial phylogeny. Evolutionary processes such as incomplete lineage sorting and introgression can cause gene tree discordance and complicate phylogenetic inferences, but are not accounted for in phylogenetic analyses of concatenated data. We generated a high-resolution data set of autosomal introns from several individuals per species and of Y-chromosomal markers. Incorporating intraspecific variability in coalescence-based phylogenetic and gene flow estimation approaches, we traced the genealogical history of individual alleles. Considerable heterogeneity among nuclear loci and discordance between nuclear and mitochondrial phylogenies were found. A species tree with divergence time estimates indicated that ursine bears diversified within less than 2 My. Consistent with a complex branching order within a clade of Asian bear species, we identified unidirectional gene flow from Asian black into sloth bears. Moreover, gene flow detected from brown into American black bears can explain the conflicting placement of the American black bear in mitochondrial and nuclear phylogenies. These results highlight that both incomplete lineage sorting and introgression are prominent evolutionary forces even on time scales up to several million years. Complex evolutionary patterns are not adequately captured by strictly bifurcating models, and can only be fully understood when analyzing multiple independently inherited loci in a coalescence framework. Phylogenetic incongruence among gene trees hence needs to be recognized as a biologically meaningful signal. |
|
|
|
Post by brobear on Jan 5, 2021 5:06:59 GMT -5
www.sciencedaily.com/releases/2017/04/170419093151.htm Bears breed across species borders Date: April 19, 2017 Source: Senckenberg Research Institute and Natural History Museum Summary: Scientists have sequenced the entire genomes of four bear species, making it now possible to analyze the evolutionary history of all bears at the genome level. It shows that gene flow, or gene exchange, between species by extensive hybridization, is possible between most bear species, not only polar and brown bear. The DNA samples of different bear species came from different European zoos, underlining their importance not only for conservation, but also for research. The study also questions the existing species concept in general, because other genome studies have frequently found gene flow among species. Senckenberg scientists have sequenced the entire genomes of four bear species, making it now possible to analyze the evolutionary history of all bears at the genome level. It shows that gene flow, or gene exchange, between species by extensive hybridization, is possible between most bear species -- not only polar and brown bear. The DNA samples of different bear species came from different European zoos, underlining their importance not only for conservation, but also for research. The study published in Scientific Reports also questions the existing species concept in general, because other genome studies too have, frequently found gene flow among species. Pizzly, grolars or "capuccino bears" are common names of the offspring resulting from the mating of grizzly bears (Ursus arctos) and polar bears (Ursus maritimus). "Such hybrids among bears are not as rare as we have hitherto assumed," says Prof. Dr. Axel Janke of the Senckenberg Biodiversity and Climate Research Center in Frankfurt. In a large-scale analysis, a team of scientists led by the German evolutionary geneticist has sequenced six complete bear genomes. Each genome is about 2.5 billion base pairs large. "With these new data of the sun bear, sloth bear, Asiatic black bear and spectacled bear, we now have the genomes of all known bear species," adds Janke.
|
|
|
|
Post by brobear on Jan 5, 2021 5:10:22 GMT -5
www.upi.com/Science_News/2017/04/19/Scientists-find-evidence-of-gene-flow-across-most-bear-species/6711492617374/ April 19 (UPI) -- New genomic research suggests mosts bears breed across species boundaries and are capable of producing fertile hybrids. Scientists have long assumed species hybridization is unsustainable and typically yields infertile offspring. For example, a mule, the offspring of a horse and donkey, can't reproduce. Polar-grizzly hybrids -- often called grolars, pizzly bears or "cappuccino bears" -- are often fertile. Recently, scientists from the Senckenberg Research Institute and Natural History Museum in Germany completed the genomic sequencing of four new bear species. "With these new data of the sun bear, sloth bear, Asiatic black bear and spectacled bear, we now have the genomes of all known bear species," Axel Janke, a scientist at the Senckenberg Biodiversity and Climate Research Center in Frankfurt, said in a news release. The results -- detailed in the journal Scientific Reports -- revealed evidence of interspecies gene flow, or gene exchange, throughout ursine evolution. Several studies have suggested bear hybridization is the product of climate change -- warmer temperatures push brown bears northward into polar bear territory. But the new research suggests bear hybridization has been prevalent for thousands of years. The research also showed gene flow between polar and sun bears, despite significant geographical separation between the two species. Only an intermediary host, like brown-polar hybrids, can explain the shared genetics. "By hybridization the brown bear could pass these polar bear genes on to other bear species in Asia," Janke said. Reproductive exclusivity or isolation have long been used as a defining marker of special distinction, but the latest research calls such logic into question. "We have to ask ourselves: Does the species concept still hold true, given there is evidence of gene flow not only in bears, but also in other animals?" Janke said. "Therefore, what do we need to protect for the future -- species or genomic diversity?" Janke believes the answer is genomic diversity. "What we must preserve... is genetic variation to protect diversity and to allow adaptation to future environmental changes," he said.
|
|
|
|
Post by King Kodiak on Jan 5, 2021 6:00:51 GMT -5
They basically just talk about bear evolution and how they figured that out using the gene flow across different species. Sometimes they wonder if they are even different species.
|
|
|
|
Post by brobear on Jan 5, 2021 8:09:56 GMT -5
They basically just talk about bear evolution and how they figured that out using the gene flow across different species. Sometimes they wonder if they are even different species. This I understand. But can you explain Nuclear DNA Y chromosome mtDNA gene flow scenarios of (A) Cloudogram of species trees from *BEAST analysis, based on 14 autosomal introns and 5.9 kb of Y-chromosomal sequence (90,000 species trees)? |
|
|
|
Post by King Kodiak on Jan 5, 2021 10:31:44 GMT -5
They basically just talk about bear evolution and how they figured that out using the gene flow across different species. Sometimes they wonder if they are even different species. This I understand. But can you explain Nuclear DNA Y chromosome mtDNA gene flow scenarios of (A) Cloudogram of species trees from *BEAST analysis, based on 14 autosomal introns and 5.9 kb of Y-chromosomal sequence (90,000 species trees)? |
|
|
|
Post by King Kodiak on Jan 7, 2021 14:45:41 GMT -5
brobear Looks like "Beast" is some kind of software for analysis of molecular sequences related by an evolutionary tree:
BEAST: Bayesian evolutionary analysis by sampling trees
Abstract
Background
The evolutionary analysis of molecular sequence variation is a statistical enterprise. This is reflected in the increased use of probabilistic models for phylogenetic inference, multiple sequence alignment, and molecular population genetics. Here we present BEAST: a fast, flexible software architecture for Bayesian analysis of molecular sequences related by an evolutionary tree. A large number of popular stochastic models of sequence evolution are provided and tree-based models suitable for both within- and between-species sequence data are implemented.
bmcecolevol.biomedcentral.com/articles/10.1186/1471-2148-7-214
|
|
|
|
Post by brobear on Aug 12, 2021 8:53:38 GMT -5
*I have to admit it. I personally view the Ailuropoda, such as our living giant panda, as being almost a bear.
Same as with the Amphicyonidae ( bear dogs ) and the Hemicyonidae ( dog bears ).

|
|
|
|
Post by brobear on Aug 13, 2021 0:35:47 GMT -5
Almost a Bear:
1- Agriarctos depereti.
2- Agriarctos vighi.
3- Agriarctos gaali.
4- Ailurarctos yuanmouensis.
5- Ailurarctos lufengensis.
6- Kretzoiarctos beatrix.
7- Ailuropoda microta.
8- Ailuropoda wulingshanensis.
9- Ailuropoda minor.
10- Ailuropoda melanoleuca - giant panda.
11- Ailuropoda qinlingensis - giant panda.
|
|
|
|
Post by brobear on Sept 24, 2022 6:38:39 GMT -5
Bears are giant weasels pterosaurheresies.wordpress.com/2018/09/02/bears-are-giant-weasels/ This post and the two previous ones here and here were prompted by a YouTube video on Bear-dogs.  Today we’ll talk about bears and their place in the cladogram of the Carnivora. Turns out they are not related to dogs (contra Matthews 1930, Wang 1994), so there should be no such thing as a bear-dog (contra Tomiya and Tseng 2016). Taxon exclusion, as usual, is the problem with these earlier, more focused studies. |
|
|
|
Post by brobear on Sept 24, 2022 6:42:14 GMT -5
DNA has it backwards. Flynn et al. 2005 recovered extant Carnivora relationships, but nested cats and dogs at the base of the clade (Fig. 2). Count the toes. Basal taxa, like weasels, have five toes in contact with the substrate. And which taxa look most like the outgroup arboreal didelphids? Civets and weasels. And where are the civets? They were not included. Ironically, this is an example of DNA matching trait analysis, except Flynn et al. reversed the order, had the cladogram upside-down. Of course, the lack of fossils obscures more precise relationships and omits many key extinct taxa. We looked at problems with DNA cladograms here, here and here.  |
|
|
|
Post by brobear on Sept 24, 2022 6:45:31 GMT -5
By contrast the large reptile tree (LRT, 1278 taxa; subset Fig. 3) nest cats and dogs as the most highly derived members of the Carnivora. They only have four toes in contact with the substrate and they look different than outgroup didelphids. And they are highly arboreal. Civets (genus: Nandinia) and vulpavids (genus: Vulpavus) nest at the base of the Carnivora in the LRT (Fig. 3). Civet-like arboreal didelphids (genus: Caluromys) are outgroups. Bears, like Ursus maritimus (= polar bear; Fig. 4) nest between the weasels (genera: Mustela and Puijila) and the giant weasels (genus: Stylinodon) with teeth that continued growing throughout its lifetime and giant canines that came to resemble rodent-like incisors. The aquatic forms, Palaeonsinopa and seals like Phoca are bear cousins.  |
|
|
|
Post by brobear on Sept 24, 2022 6:48:11 GMT -5
Puijila (Rybczynski et al., 2009) was originally considered “a walking seal”, but here links weasels to bears and weasels to seals (but not sea lions, contra Rybczynski et al., 2009, because the Pinnipedia is diphyletic). Remember: (Don’t trust teeth alone because they change with diet. Always score traits for the entire skull and body whenever possible to get the closest possible relatives from a long list of candidates).  |
|
|
|
Post by brobear on Sept 24, 2022 6:50:16 GMT -5
Earlier we looked at another pair of giant weasels, now also allied to bears, the traditionally enigmatic Stylinodon (Fig. 5) and Psittacotherium. Let’s talk about the traditional origin of bears. There is a mink-sized early Oligocene taxon known from the front half of a skull, Amphicynodon (Fig. 5; Wang , McKenna and Dashzeveg 2005) that has been the traditional ancestor of bears. In the LRT Amphicynodon nests close to bears, but closer to Stylinodon and Psittacotherium, taxa never tested with Amphicynodon before.  |
|
|
|
Post by brobear on Sept 24, 2022 8:16:35 GMT -5
|
|
|
|
Post by brobear on Sept 24, 2022 14:45:10 GMT -5
So Mustelidae and Mephitidae (weasels and skunks), Procyonidae (raccoons), and Ailuridae (red panda) are all close relatives of bears as well as Phocidae ("earless" seals), Odobenidae (walrus), and (possibly) the Otariidae ("eared" seals).
|
|